HELM-BERT: Topology-aware representations for chemically modified peptides
Abstract
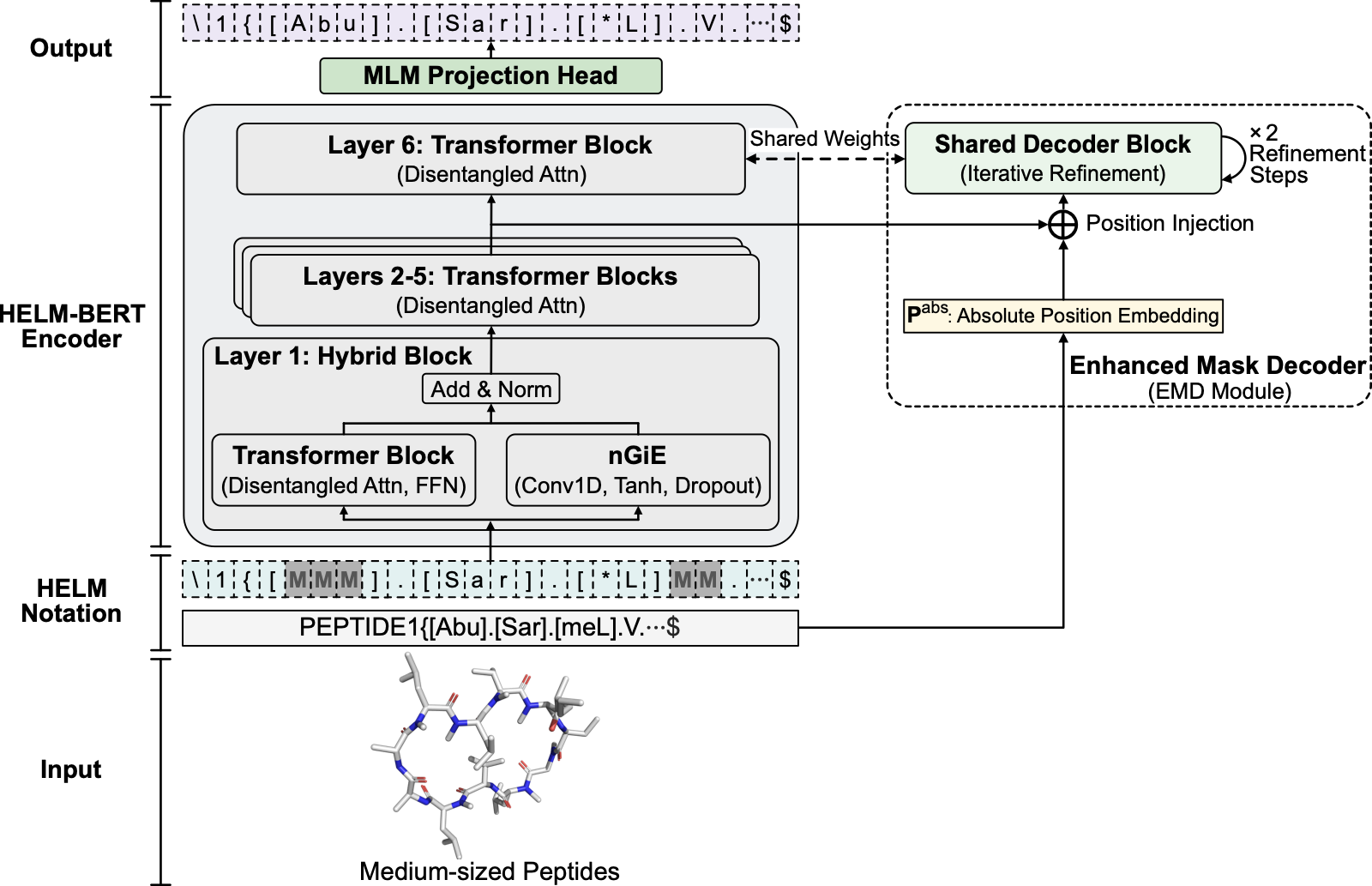
Chemically modified and macrocyclic peptides are increasingly important therapeutics, yet current molecular representation models do not natively represent chemical modification and covalent topology in a unified way. Atom-level strings obscure macrocyclic connectivity, whereas protein sequence models cannot encode non-canonical residues and explicit crosslinks. Here we pretrain an encoder-only transformer directly on Hierarchical Editing Language for Macromolecules (HELM) notation, which specifies monomer identity and connectivity. In this work, we show that the resulting representations achieve top mean performance in cyclic peptide membrane permeability prediction (random split R2 = 0.668; retaining top mean performance under a Murcko scaffold split), exceeding external pretrained SMILES-based encoders. An architecture-matched SMILES control narrowed the HELM–SMILES gap under full finetuning, whereas HELM-BERT retained clearer advantages in frozen-representation settings. HELM-BERT also preserves HELM-specified macrocyclic topology in a linearly accessible form and supports competitive peptide–protein interaction prediction across complementary Propedia and ChEMBL benchmarks. More broadly, these findings suggest that pretraining directly on notation that makes structural constraints explicit offers a transferable strategy for biomolecular modalities that fall between small-molecule chemistry and protein sequence.
Accepted 7 May 2026; DOI to be assigned upon publication.
